In our working group, we have a diverse array of database related projects.
This work is conducted by Lars Langer (supervised by Prof. Christian Wirth, our working group, and Prof. Manuel Burghardt, Digital Humanities)
Nature’s non-material contributions to people are difficult to quantify. One aspect of them, nature’s contribution to communication, has so far been neglected. Recent advances in automated language processing tools enable us to quantify biodiversity patterns underlying the distribution of plant and animal taxon labels in creative literature which we term BiL (biodiversity in literature). We assume BiL to provide a proxy for people’s openness to NCCs.
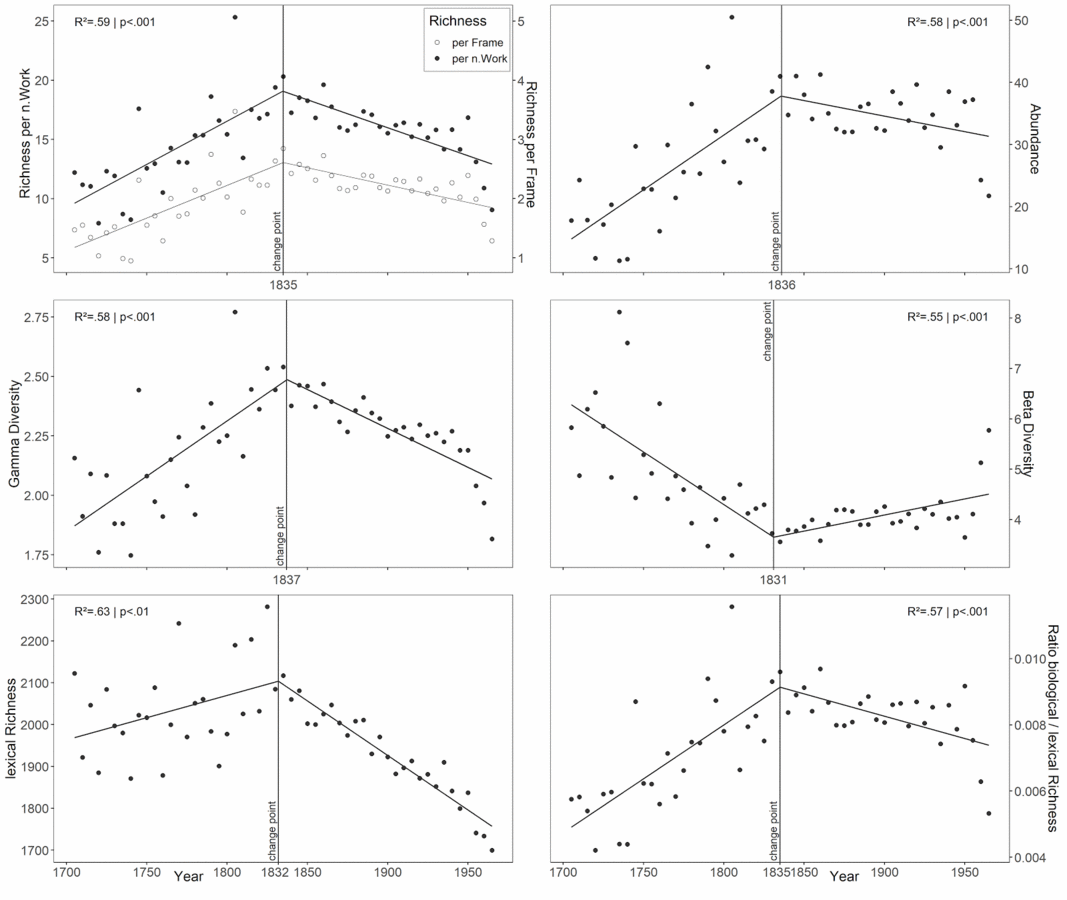
We assembled a comprehensive list of 240,000 English biological taxon labels. We pre-processed and searched a subcorpus of digitized literature of Project Gutenberg for these labels. We quantified changes of biodiversity indices commonly used in ecological studies for 16,000 books of 4,000 authors as proxies for BiL between 1705 and 1969. We additionally relate our findings to the general lexical richness of the language as represented in our corpus. Have a look at our newest results:
We investigate possible causes of the resulting patterns of BiL, to which end we assemble and assess large amounts of metadata on our works, authors as well as listed animals and plants.
sROOT was initiated by Alexandra Weigelt and Liesje Mommer in 2018 as a working group supported by sDiv, the Synthesis Centre of the German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig.
During the first meeting in October 2018 the group assembled the largest global database of fine root traits including measurements of four key functional traits (specific root length, root diameter, root tissue density, and root nitrogen concentration) on > 1700 species. This database (GrooT) was then used to conceptualize and test a new framework on root trait correlation, the root economics space. This work resulted two papers which were finalized and submitted after the second workshop in May 2019, one database paper led by Nathaly Guerrero-Ramirez (Guerrero-Ramirez, Mommer, sROOTies & Weigelt, Global Ecology and Biogeography 2020) and the paper on the root economics space led by Joana Bergmann (Bergmann, Weigelt, sROOTies & Mommer, Science Advances 2020). In addition, during the second workshop, we started working on a third paper combining GrooT with the global plant database sPLOT to ask if and how root traits define species compositions along global environmental gradients. This paper was led by Daniel Laughlin and was finalized in the third workshop in January 2020 and after (Laughlin, Mommer, …, Weigelt, Nature Ecology and Evolution 2021). The fourth ‘sROOTies’ paper was also initiated during the third workshop and studies trait correlations on the whole plant basis linking the leaf economics spectrum and the root economics space. This paper was finalized by a core team of sROOT members, Karl Andraczek, Colleen Iverson, Joana Bergmann, Luke McCormack, together with the two leading authors Alexandra Weigelt and Liesje Mommer. This paper is currently under revision in New Phytologist.
Further information in the sDiv working group
PhenObs is performed by Dr. Martin Freiberg.
Founded in 2017, PhenObs is a global network of Botanical Gardens for phenological observations. PhenObs unites scientists, students and citizen scientists in answering the question, what kind of influence climate change has on the phenology of herbaceous plants. The phenology and functional plant traits are measured following standardized procedures. Botanical gardens are ideal for this, as they provide an excellent foundation through their species richness to investigate many species of different habitats. Phenology itself describes the chronological sequence of biological development successions. For plants these are, e.g., foliation, flower and fruit development and senescence.
Climate change shifts these developments. E.g. within the last 100 years, the onset of spring started 11 days earlier. This influences biodiversity, biotic interaction and ecosystem services.The aim of PhenObs is to better understand the phenological shifts in herbaceous plants through abiotic and biotic changes. Moreover we would like to investigate whether those shifts could be predicted.
LCVP und LifeGate is performed by Dr. Martin Freiberg.
Some of the fundamental questions in biology are still unanswered: How many species are there? Do more species go extinct than new ones are discovered? Where are the species hotspots – and why? Where is the largest scientific demand? Extensive analyses of all kinds of resources are necessary to answer these questions, because already described species should not be described again.
To tackle this problem a substantial databank of 1.3 Mio botanical names (“LCVP”, the Leipzig Catalogue of Vascular Plants”) has been built up since 2012. It is continuously updated with the most recent publications and so far refers to about 360.000 accepted vascular species. LCVP as well as other data banks on further phyla of life are established to include their phylogenetic relationships. This is the underlying basis for a new webpage (lifegate.idiv.de) which will be launched in August 2021. LifeGate facilitates a visual and interactive gate to the diversity of life and is globally already supported by more than 1000 photographers.
Conducted by Dr. Verónika Ceballos N. (supervised by Prof. Christian Wirth, University Leipzig, Assist. Prof. Kathryn Barry, Utrecht University, and Dr. Nadja Rüger, IDiv)
Long title: “How biodiversity change alters the biotic context of biodiversity-ecosystem functioning relationships - a synthesis of models, experiments and observations”
It has been predicted that the change in global biodiversity (a pressing issue of our time) will likely alter the way ecosystems function. The ecosystem-functioning dependence on biodiversity is driven by mechanisms that will be explored in this project.
This project is rooted in the SimNet model intercomparison project, which is a collaborative group of community modelers funded by the German Centre for Integrative Biodiversity Research (iDiv). It consists of three work packages that incorporate synthesis of theory, empirical data from experiments, and observational data at the global scale
Conducted by Dr. David Schellenberger Costa (supervised by Prof. Christian Wirth, University Leipzig)
PlantHub is a project created by the German Centre of Integrative Biodiversity Research (IDiv) and partners to increase exposure of data collected by individual researchers and projects as well as research associated to this data. Research groups at idiv work on different biodiversity-related topics. Data collected ranges from environmental variables in high spatial and temporal resolution over phylogenetic information on plants and animals to vegetation surveys done all over the planet. This wealth of information is used for research, but it lacks the visibility to be known even by all members of idiv itself. For external researchers or the general public, the barriers for accessing the data are even higher. Through PlantHub, we want to offer a single “entry point” to the idiv biodiversity data universe, with a range of options to visualize data, learn about individual projects, find useful workflows for data harmonization and analysis, and read or redo our newest research. For researchers, this could lead to creating new hypotheses, gaining access to data needed for analyses, or checking species lists for consistency. For the public, PlantHub could be a source of information on biodiversity in general, but also to new worthy travel locations or just beautiful plants to plant in one’s garden. Currently, projects involved with PlantHub are TRY, a plant trait data base, sPlot, a vegetation survey data base, sMon, a research project on biodiversity trends in Germany, PhenObs, a project on plant phenology in botanical gardens, LifeGate and LCVP, data bases on plant phylogeny and taxonomy, and the dataCube, a collection of environmental data in space and time.
For further information, please contact David Schellenberger Costa.